This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
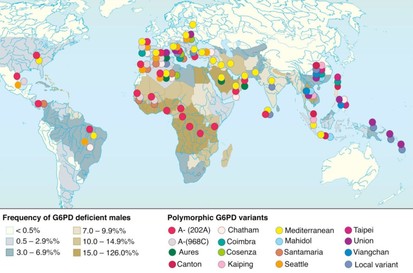
G6PD Deficiency (G6PDd) is the most prevalent enzymopathy in the world, affecting over 400 million people worldwide. The wide distribution of the deficiency is due to the fact that nearly 200 different single nucleotide polymorphisms can cause the enzyme to be deficient [1]. G6PDd is most commonly known for causing neonatal jaundice (skin yellowing, neurological deficiencies) and hemolytic anemia (low blood pressure, skin and eye yellowing, weakness, heart disease) [2].
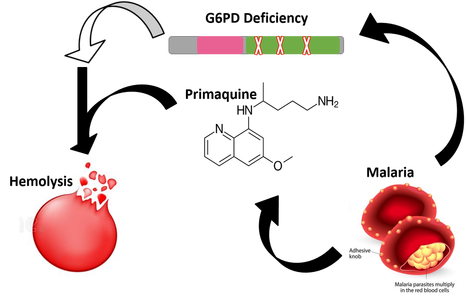
G6PD deficiency itself does not cause hemolysis, it just makes one more susceptible to reactive oxygen species, as the inability to produce NADPH weakens the cells. For hemolysis to actually be triggered, there needs to be an external oxidizing factor. One of the most common external triggers is the antimalarial Primaquine [3]. Primaquine's prevalence in triggering hemolysis is due to the balancing selection between G6PD and Malaria. Malaria endemicity selects for G6PD deficient individuals in the population, because those with healthy red blood cells are more likely targets of malarial parasites. However, for those with G6PD deficiency, treatment with Primaquine can be fatal. G6PD deficiency-induced hemolysis triggered by Primaquine is a major obstacle in malaria eradication. One way to overcome this obstacle is to discover a way to increase NADPH levels in G6PD deficient individuals so they can be safely prescribed the antimalarial.
|
G6PD Deficiency weakens the cell through the decreased ability to produce NADPH, but does not directly cause hemolysis External oxidizing triggers, such as Primaquine, cause hemolysis in G6PD deficient individuals |
Malaria causes G6PDd to be more prevalent in a population because weakened blood cells confer some protection against infection by malarial parasites
Primaquine is an antimalarial, and the only antimalarial that is effective enough to prevent relapse. |
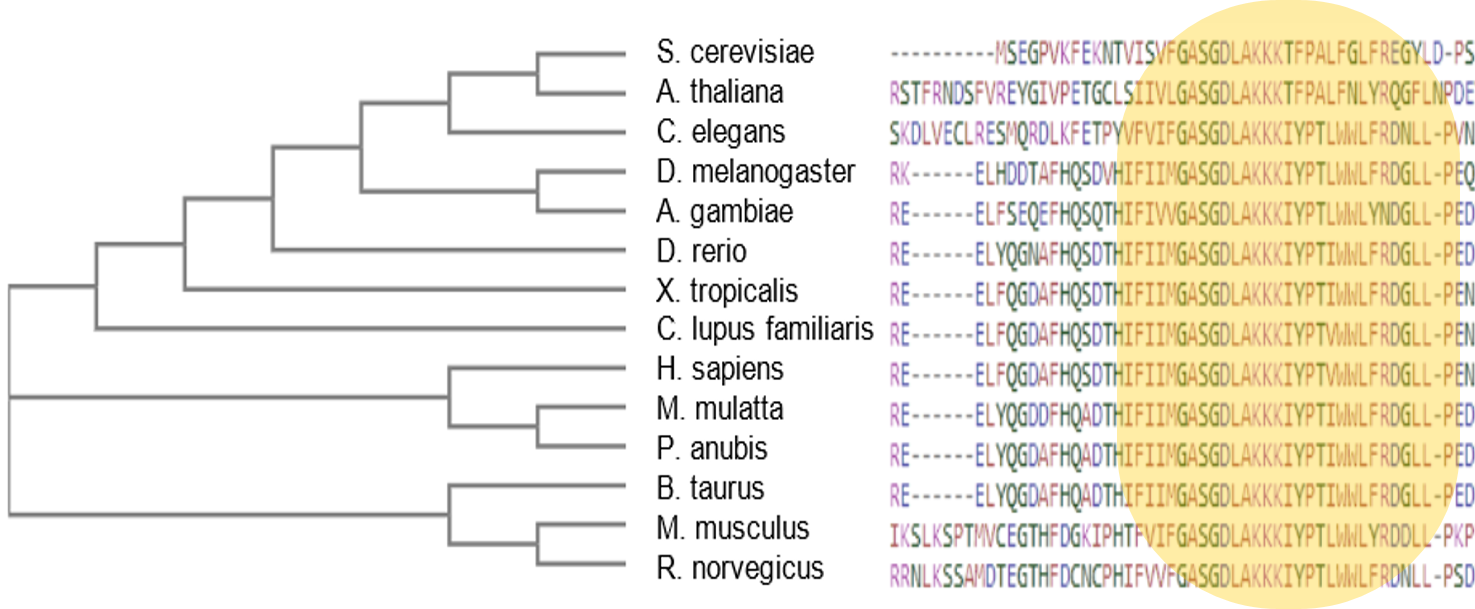
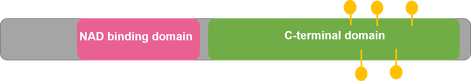
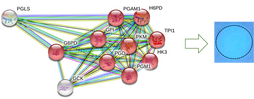
The G6PD enzyme can be found throughout the cell cytoplasm [4]. This makes sense, as its primary molecular functions and biological processes are involved in NAD binding and NADPH production, respectively. The G6PD protein interaction network further demonstrates G6PD's role in the NADPH production pathway, as the majority of the protein interactions are with other proteins involved in NADPH metabolism. Because NADPH is a necessary reducing agent in almost all living species, G6PD is well-conserved, with both the NAD-Binding Domain and C-Terminal Domain present in organisms from Humans to C. elegans to Arabidopsis.
In 1985, researchers Lucio Luzzato and Giorgio Battistuzzi wanted to determine how methylation of the G6PD gene would impact G6PD levels, and by extension, NADPH production [5]. They found that DNA methylation is positively correlated with G6PD levels. However, this study was only done on males. My primary goal is to determine how methylation affects G6PD levels and NADPH production in pregnant females. I am interested in pregnant females because pregnancy increases the risk for anemia, lowering the blood cell count [6]. Additionally, malaria during pregnancy causes more distress than it does in non-pregnant individuals [7]. For G6PD deficient women with fewer red blood cells, hemolysis triggered by Primaquine is even more deadly. My hypothesis is that G6PD expression and NADPH production in pregnant women can be upregulated through methylation of the G6PD gene.
|
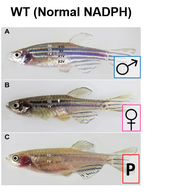
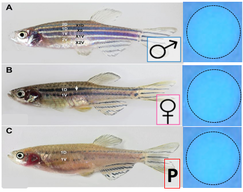
Though the G6PD gene is well-conserved in many model organisms, I will be using zebrafish as a model organism to conduct my experiment. G6PD deficient zebrafish and humans share similar phenotypes when triggered by external factors, and hemolysis is easy to observe in zebrafish due to their transparent nature [8]. In addition, zebrafish are relatively inexpensive with shorter generation times, compared to other common model organisms.
Throughout my study, I will be comparing the methylation patterns and NADPH levels of male, nonpregnant female, and pregnant (egg-producing) zebrafish. |
Aim 1: Knockout conserved methylation sites of G6PD to identify methylation sites associated with normal levels of NADPH production in pregnant females.
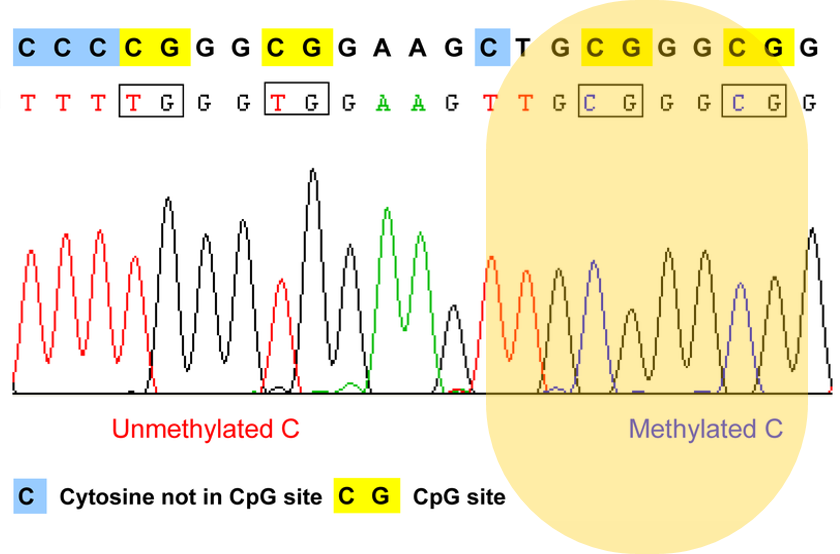
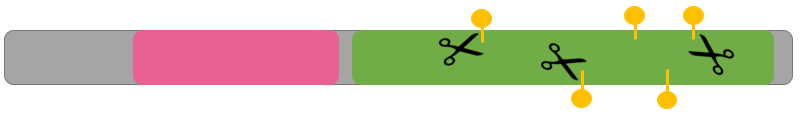
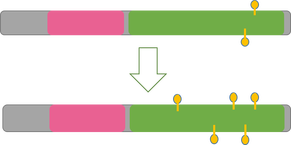
In the first stage of Aim 1, I will first use high-throughput bisulfite sequencing to determine the methylation pattern of wild-type zebrafish of all three groups. I will then use Clustal-Omega to align the sequences of homologs to find the most well-conserved methylation sites. My rationale for Aim 1 is that it is important to determine if methylation has the same upregulating effect in pregnant zebrafish as it does in males.
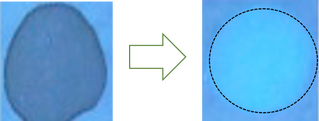
In the second stage of Aim 1, I will use the fluorescent spot test to determine the NADPH levels associated with normal WT zebrafish with a normal methylation pattern. NADPH is naturally fluorescent, and the fluorescent spot test is a qualitative method to measure NADPH production based on brightness. I will then use CRISPR/Cas-9 to knockout the most conserved methylation sites found in stage 1. After knocking out the most conserved methylation sites, I will conduct the fluorescent spot test once again to determine the changes in NADPH levels due to the altered methylation pattern.
|
Initial Fluorescent Spot Test followed by CRISPR/CAS9 to knockout well conserved methylation.
|
Hypothesized result of knockout
|
I hypothesize that there will be a specific and conserved methylation pattern of the G6PD gene associated with normal G6PD function (NADPH production) and that knockout of these sites will result in lower NADPH levels.
Aim 2: Identify a small molecule that affects methylation of G6PD and can be taken with Primaquine in pregnant females without hemolysis
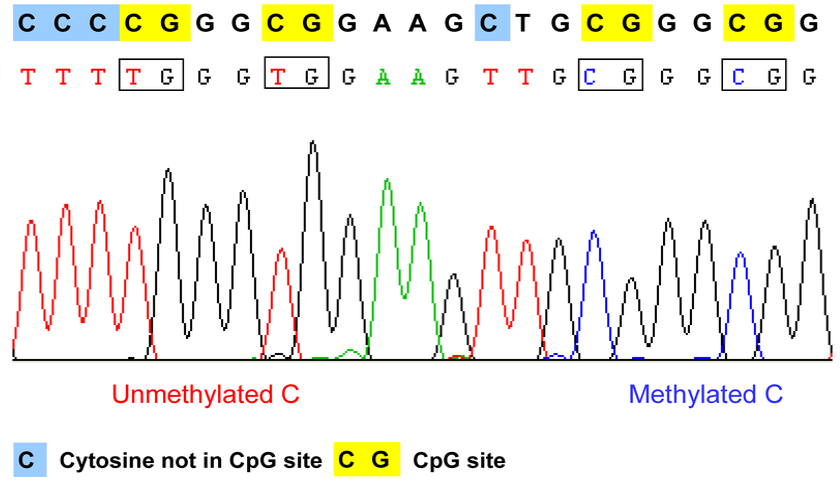
In the first stage of Aim 2, I will conduct bisfulfite sequencing of the pregnant, G6PD deficient zebrafish. I predict there will be fewer methylated sites.
|
Bisulfite Sequencing of Pregnant, G6PD deficient zebrafish
|
Hypothesized result
|
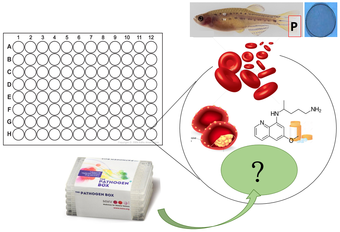
In the second stage of Aim 2, I will conduct a chemical screen to identify a small molecule that affects methylation, destroys malaria, and can be taken with Primaquine without causing hemolysis in G6PD deficient, pregnant zebrafish. I will be testing small molecules from the Pathogen Box, made by Medicines for Malaria Ventures.
I hypothesize that there will be a small molecule that modifies G6PD methylation in pregnant females, resulting in stable and sufficient NADPH levels when Primaquine is administered.
Aim 3: Determine changes in G6PD Protein Interaction Network caused by methylation changes
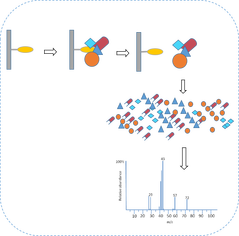
In stage 2 of Aim 3, I will conduct tandem-affinity-purification followed by mass spectrometry to study the protein interaction of the control and treatment groups. My rationale for assessing the protein interactions is to determine if they mechanism by which methylation affects NADPH production is by helping regulate the protein complexes G6PD forms with other proteins involved in NADP metabolism.
|
TAP-MS of control (methylated) and treatment (unmethylated) groups
|
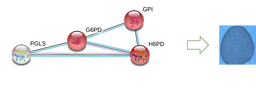
Hypothesized Results
|
I hypothesize that methylation levels affect NADPH levels through the regulation of protein-interaction networks, and that when methylation is knocked-out, there will be fewer protein interactions between G6PD and other NADP metabolism controlling proteins.
Future Directions
|
G6PD deficiency is currently a major obstacle for malaria eradication. One way this could be overcome is through pharmacogenomics. Pharmacogenomics is assessing an individual's genes to determine their likely drug response, and treat accordingly, with different drugs or dosages for the same condition, depending on one's genetics [9]. The chemical screen conducted in Aim 2 would be the perfect launchpad to begin creating different drugs that could be taken in tandem with, or instead of, Primaquine for G6PD deficient individuals who contract malaria.
Additionally, if I had to add a fourth aim, I would have like to look at how the post-translational modifications may change depending on methylation levels, and whether that would have an effect on level of NADPH production. Namely, I would have looked at phosphorylation sites, as they are the most common functional group that alters protein folding.
|
Rough Draft 1
| xue_gen564talk.pptm | |
| File Size: | 2516 kb |
| File Type: | pptm |
Rough Draft 2
| xue_gen564talk.pptm | |
| File Size: | 2814 kb |
| File Type: | pptm |
Final Talk
| xuegen564finaltalk.pdf | |
| File Size: | 2317 kb |
| File Type: | |
References
[1] Cappellini M. (2008). "Glucose-6-phosphate dehydrogenase deficiency". The Lancet 371: 64-74
[2] Luzzatto L. (2006). "Glucose 6-phosphate Dehydrogenous Deficiency: From Genotype To Phenotype". Journal of European Hematology Association 91: 1303-
1306
[3] Baird K. (2015). “Origins and Implications of Neglect of G6PD deficiency and primaquine toxicity in Plasmodium vivax malaria”. Pathogens and Global
Health 109(3): 93-106.
[4] String. (2017). "G6PD." Retrieved from string-db.org/cgi/network.pl?taskId=fghKTuOaAdTx
[5] Battistuzzi G. (1985). “Tissue-specific levels of human glucose-6-phosphate dehydrogenase correlate with methylation of specific sites at the 3’ end of
the gene.” Proceedings of the National Academy of Sciences:Genetics 82: 1465-1469.
[6] American Society of Hematology. (2017). "Anemia and Pregnancy." Retrieved from http://www.hematology.org/Patients/Anemia/Pregnancy.aspx
[7] Schantz-Dunn J. (2009). "Malaria and Pregnancy: A Global Health Perspective." Reviews in Obstetrics and Gynecology 2(3): 186–192.
[8] Patrinostro X. (2013). "A model of glucose-6-phosphate dehydrogenase deficiency in the zebrafish." Experimental Hematology
[9] Genetics Home Reference (2018). "What is Pharmacogenetics?" Retrieved from https://ghr.nlm.nih.gov/primer/genomicresearch/pharmacogenomics
[1] Cappellini M. (2008). "Glucose-6-phosphate dehydrogenase deficiency". The Lancet 371: 64-74
[2] Luzzatto L. (2006). "Glucose 6-phosphate Dehydrogenous Deficiency: From Genotype To Phenotype". Journal of European Hematology Association 91: 1303-
1306
[3] Baird K. (2015). “Origins and Implications of Neglect of G6PD deficiency and primaquine toxicity in Plasmodium vivax malaria”. Pathogens and Global
Health 109(3): 93-106.
[4] String. (2017). "G6PD." Retrieved from string-db.org/cgi/network.pl?taskId=fghKTuOaAdTx
[5] Battistuzzi G. (1985). “Tissue-specific levels of human glucose-6-phosphate dehydrogenase correlate with methylation of specific sites at the 3’ end of
the gene.” Proceedings of the National Academy of Sciences:Genetics 82: 1465-1469.
[6] American Society of Hematology. (2017). "Anemia and Pregnancy." Retrieved from http://www.hematology.org/Patients/Anemia/Pregnancy.aspx
[7] Schantz-Dunn J. (2009). "Malaria and Pregnancy: A Global Health Perspective." Reviews in Obstetrics and Gynecology 2(3): 186–192.
[8] Patrinostro X. (2013). "A model of glucose-6-phosphate dehydrogenase deficiency in the zebrafish." Experimental Hematology
[9] Genetics Home Reference (2018). "What is Pharmacogenetics?" Retrieved from https://ghr.nlm.nih.gov/primer/genomicresearch/pharmacogenomics