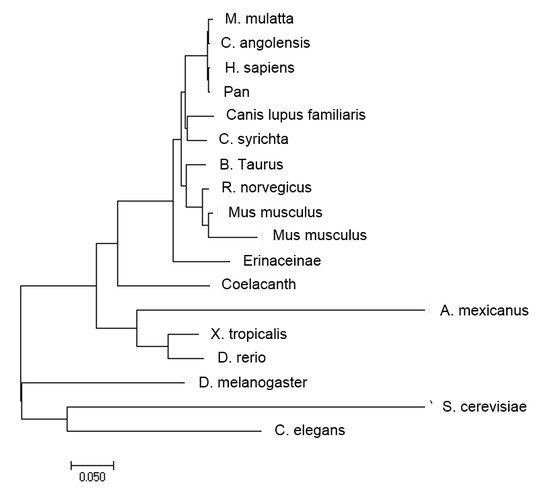
Phylogeny
This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
What is Phylogeny?
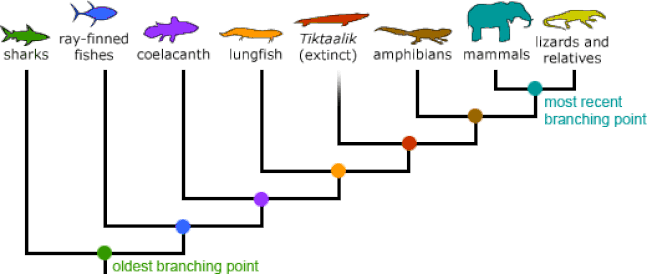
Phylogeny is the study of the evolutionary relationship of organisms [1]. Phylogenies do not represent progress of organisms, as once believed, but rather branching points in evolutionary history of speciation events. All currently living organisms are equally "evolved" [2]. There are a variety of methods that can be used to build phylogenies. The most common are maximum likelihood and neighbor-joining.
Building Phylogenies
1. Obtain Sequences in FASTA format (save as .txt file)
2. Align Sequences using ClustalOmega
3. Create Simple Phylogeny in ClustalOmega Using Neighbor-joining Method
4. Create Trees in MEGA
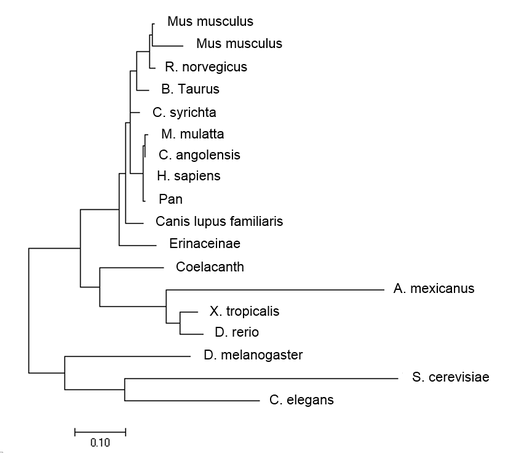
Maximum Likelihood:
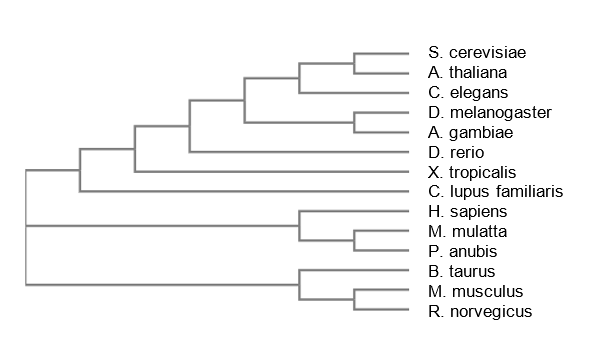
Neighbor-joining
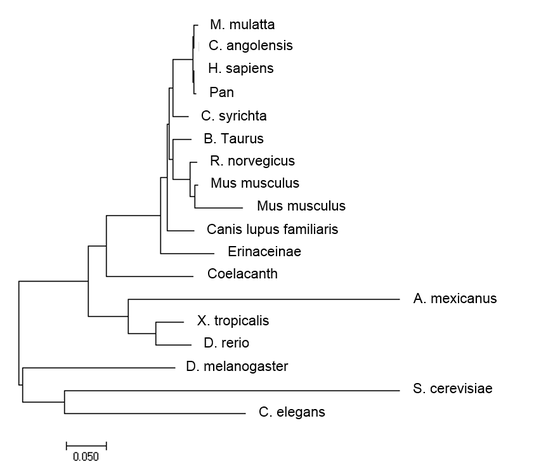
Minimum Evolution:
What is the Difference between Neighbor-joining, Maximum Likelihood, and Average Distance methods?
Maximum Likelihood first uses another method such as distance measurement or neighbor-joining, then adjusts the branch lengths depending on this likelihood of the observed branching [3]. Neighbor-joining uses BLOSSUM similarity score to determine which species are most closely related, and joins the most related species together. The branch lengths are dependent on the number of changes that have occurred since the splitting event. Average Distance method is based on the evolutionary distance of each species from one another. Unlike Maximum Likelihood and Neighbor-joining, Each branch is equidistant from its most recent common ancestor.
Discussion
Phylogenies help us visualize that the G6PD gene and gene product are highly conserved throughout plant and animal species. It plays a vital role in almost all cells. Furthermore, phylogenetcs also allows us to compare how the gene and protein are similar or different among homologs.
Discussion
Phylogenies help us visualize that the G6PD gene and gene product are highly conserved throughout plant and animal species. It plays a vital role in almost all cells. Furthermore, phylogenetcs also allows us to compare how the gene and protein are similar or different among homologs.
References
[1]. University of California-Berkeley. (2008). Phylogenetics. Retrieved from http://evolution.berkely.edu/evolibrary/article/phylogenetics_02
[2]. University of California-Berkeley. (2008). Misconceptions about Evolution. Retrieved from https://evolution.berkeley.edu/evolibrary/misconceptions_teacherfaq.php
[3]. Singh, M. (2003). Phylogenetics. Retrieved from https://www.cs.princeton.edu/~mona/Lecture/phylogeny-slides.pdf
[1]. University of California-Berkeley. (2008). Phylogenetics. Retrieved from http://evolution.berkely.edu/evolibrary/article/phylogenetics_02
[2]. University of California-Berkeley. (2008). Misconceptions about Evolution. Retrieved from https://evolution.berkeley.edu/evolibrary/misconceptions_teacherfaq.php
[3]. Singh, M. (2003). Phylogenetics. Retrieved from https://www.cs.princeton.edu/~mona/Lecture/phylogeny-slides.pdf