Protein Interaction Networks
This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
What are protein interaction networks?
Protein Interaction Networks show all of the proteins that your protein of interest interacts with. Useful tools include the String protein-protein interaction database, which displays the protein of interest connected to all the proteins it interacts with by a line or "string." These interactions can be sorted by gene ontology terms, homology, and type of interaction [1]. Protein interaction networks can help researchers better understand what the protein does, and can also reveal new pathways that the protein is involved in.
The Human G6PD Protein Interaction Network
Protein Interaction Networks show all of the proteins that your protein of interest interacts with. Useful tools include the String protein-protein interaction database, which displays the protein of interest connected to all the proteins it interacts with by a line or "string." These interactions can be sorted by gene ontology terms, homology, and type of interaction [1]. Protein interaction networks can help researchers better understand what the protein does, and can also reveal new pathways that the protein is involved in.
The Human G6PD Protein Interaction Network
|
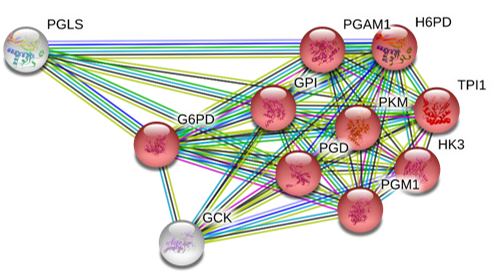
Figure 1. G6PD Protein-Protein Interactions sorted by GO term: nicotinamide nucleotide metabolic process (NADP metabolism). It makes sense that the primary protein interactions are all related to NADP metabolism, as it is the main biological process of the G6PD gene and gene products. However, this also leads us to ask what the other proteins are involved in. PGLS (6-Phosphogluconolactonase) is also involved in the Pentose Phosphate Pathway. GCK (Glucokinase) plays an important role in modulating insulin secretion[2]. Interestingly, G6PD defiency is also involved in Type II diabetes, which could explain its interaction with GCK [3].
|
Comparing Human and Zebrafish Protein Interaction Networks
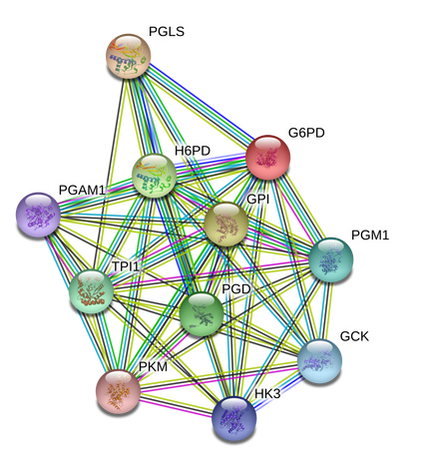
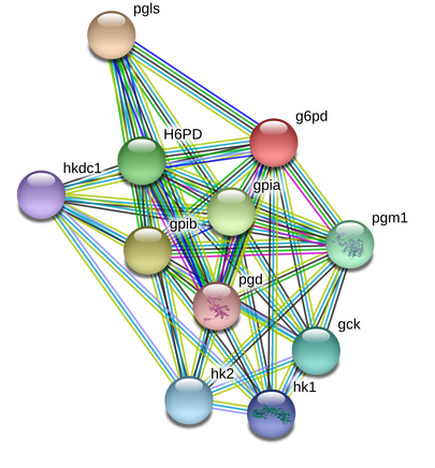
The zebrafish protein-interaction networks could not be sorted by Gene Ontology terms using String. What can be seen by comparing the two interaction networks, however, is how similar the protein interactions are.
Comparable proteins
PGLS 6-phosphogluconolactonase:.................................................................................................................................Catalyzes reaction in Pentose Phosphate Pathway
H6PD Hexose-6-phosphate dehydrogenase:................................................................Oxidizes glucose-6-phosphate and glucose, as well as other hexose-6-phosphates
GPI (gpia and gpib) Glucose-6-phosphate isomerase:.............................................................................................................................involved in gluconeogenesis
PGM1 Phosphoglucomutase1: .........................................................................................................................................................................................glucose metabolism
PGD Phosphogluconate dehydrogenase:..........................................................................................................................................................................glucose metabolism
GCK Glucokinase:............................................................Catalyzes the initial step in utilization of glucose by the beta-cell and liver at physiological glucose concentration
HK3 (hk1 and hk2) Hexokinase 3.......................................................................................................................................................................... glucose Metabolism
Human Unique Proteins
PGAM1 Phosphoglycerate mutase 1 ............................................................................2,3-bisphosphoglycerate-dependent phosphoglycerate mutase activity in the brain
TPI1 Triosephosphate isomerase:................................................................................................................................... Pentose Phosphate Pathway and Gluconeogenesis
Zebrafish Unique Proteins
hkdc1 Hexokinase domain containing 1 .............................................................................................................................................................cellular glucose homeostasis
Discussion
In both humans and zebrafish, G6PD primarily interacts with other proteins that share similar functions of glucose metabolism and NADP metabolism. Interestingly, the proteins that were unique to either humans and zebrafish were still related to glucose metabolism, and not to unique functions. I am interested in how mutations of the G6PD gene alters protein-protein interaction and other proteins that function in NAD binding.
In both humans and zebrafish, G6PD primarily interacts with other proteins that share similar functions of glucose metabolism and NADP metabolism. Interestingly, the proteins that were unique to either humans and zebrafish were still related to glucose metabolism, and not to unique functions. I am interested in how mutations of the G6PD gene alters protein-protein interaction and other proteins that function in NAD binding.
References
[1] STRING (2018). Retrieved from https://string-db.org/cgi/input.pl
[2] Uniprot GCK (2017). Retrieved from https://www.uniprot.org/uniprot/P35557
[3] Heymann A. (2012). Glucose-6-phosphate-dehydrogenase Deficiency and Type 2 Diabetes. American Diabetes Association. 35(8): e58-e58.
Figures
Header: Computational Sociology. (2018). Wikipedia Retrieved from https://en.wikipedia.org/wiki/Computational_sociology
[1] STRING (2018). Retrieved from https://string-db.org/cgi/input.pl
[2] Uniprot GCK (2017). Retrieved from https://www.uniprot.org/uniprot/P35557
[3] Heymann A. (2012). Glucose-6-phosphate-dehydrogenase Deficiency and Type 2 Diabetes. American Diabetes Association. 35(8): e58-e58.
Figures
Header: Computational Sociology. (2018). Wikipedia Retrieved from https://en.wikipedia.org/wiki/Computational_sociology